Hello! My name is Chloe Rabinowitz, and I am an undergraduate research intern through a joint collaboration between the University of Washington (UW) EarthLab and NOAA PMEL Ocean Molecular Ecology group. This summer, I sampled zooplankton from the Salish Sea and then proceeded to extract DNA and sequence species of interest. The purpose of my work has been to publish whole mitochondrial genomes to the national nucleotide database (NCBI) to enhance our understanding of zooplankton biodiversity in the region. Sequencing the full mitogenome enriches the coverage of zooplankton species by ensuring there is a reference sequence regardless of the targeted DNA amplicon. Additionally, information gained by assembling the mitochondrial genome can provide significant insight in understanding the evolutionary history of a species.
At the beginning of my internship, I worked with researchers on a Washington Ocean Acidification Center (WOAC) cruise. As we visited long-term monitored sites around the Salish Sea, I conducted vertical and oblique net tows into the ocean to a predetermined depth and pulled them up through the water to collect zooplankton samples. When I returned to land, I analyzed the zooplankton samples from previous WOAC cruises. Participating in a sampling effort allowed me to see the entire step-by-step process of my research. For the next few weeks of my internship, I worked with mentors in the Keister Lab at UW to fine-tune my zooplankton taxonomy skills. Using a microscope and my trusty forceps, I pulled out eight species of interest from the samples, seven copepods and one amphipod. Zooplankton, such as copepods and amphipods are important to the marine food web as they are the foundational food/energy source for marine invertebrates, fish, mammals, and most things in between. The health and functionality of any marine ecosystem is directly impacted by the health, diversity, and abundance of zooplankton.

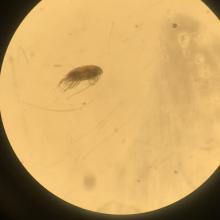
Once I identified and separated out species (even up to 2000 individuals for one species), I brought my samples to the NOAA Western Regional Office to begin my work with the PMEL ‘Omics Group. In the ‘Omics lab, I photographed each species of interest, so we had physical documentation of the specimens extracted. Then, due to their small size, we used two different extraction methodologies to determine which would result in the highest DNA yield. Next, we checked the quality and quantified how much DNA was extracted from each organism with PCR gel electrophoresis and a Qubit fluorometer. We utilized a Nanopore long-read DNA sequencer to sequence the mitochondrial genome of the amphipod species, Cyphocaris challengeri. In addition, C. challengeri and four copepod species with the highest DNA yields post-extraction, were sent for Illumina NGS sequencing. With the results of the full mitochondrial genome sequences, we will gain a better understanding of the ecology and regional impacts of this species. The information we obtain will be published to enhance scientific knowledge of their ecology and evolution, improve public awareness, and allow us to detect and monitor their presence in Puget Sound.
While zooplankton might seem like small organisms, their impact is vast. Through my work during this internship and the passion/dedication I’ve seen from my mentors, it is clear that genomics is a critical tool that allows us to understand the deep complexity of important creatures in marine ecosystems that would otherwise potentially go unnoticed.



