Ansley
Hello Ansley and others,
This works! Thanks for the help.
So to clarify: Ysequence, and the other sequences (Xsequence, Zsequence, etc), simply tell ferret that I want some numbers placed on a certain axis?
Pearse
From: Ansley C. Manke [ansley.b.manke@xxxxxxxx]
Sent: 07 May 2018 13:16
To: ferret_users@xxxxxxxx
Subject: Re: [ferret_users] putting 1D variable into 2D (lat,depth) for using in objective analysis
Hi Pearse,
Replying to the list as well -- Pearse and I looked at this a bit. The error message is
Bailing out of external function "scat2gridgauss_yz":
http://ferret.pmel.noaa.gov/Ferret/documentation/users-guide/variables-xpressions/XPRESSIONS#_SCAT2GRIDGAUSS_YZ
No data in scattered y, z points. F() data values must be defined on Y or Z axis.
**ERROR: error in external function
The arguments to the function are SCAT2GRIDGAUSS_YZ(YPTS, ZPTS, F, YAXPTS, ZAXPTS, ...) where ypts and zpts are simple lists of points, on any axis. The third argument, F, can also depend on other dimensions. This would allow for you to have a set of locations ypts, zpts, and F as a measurement at a set of times and/or a set of longitudes. This means that for the third argument, the scattered-data direction must lie along a y or z axis. The function then keeps any dependence on an X, T, E, or F axis, and finds the "scattered" data along a Y or Z direction. In the documentation,
For this case, all you need to do is recast the data for the 3rd argument onto a Y or Z axis,F-data: 3rd component of scattered input triples. This is the variable we are putting onto the grid. It must have the scattered-data direction along a Y or Z axis. May be function of X or time.
yes? define axis/y=-65:25:1/units=degrees yax-Ansley
yes? define axis/z=0:6000:50/units=meters/depth zax
yes? let xscale = 0.5
yes? let yscale = 25
yes? let cutoff = 3
yes? let no3_ind_on_y = YSEQUENCE(no3_ind)
yes? let no3int_ind = scat2gridgauss_yz(lat_ind, dep_ind, no3_ind_on_y, y[gy=yax], z[gz=zax], xscale, yscale, cutoff, 0)
yes? shade no3int_ind
On 5/4/2018 10:05 AM, Ansley C. Manke wrote:
Hi Pearse,
For your data, the arguments of scat2gridgauss_yz would just be the simple 1-D lists of data, and the result will be on your y-z grid defined by the axes yax and zax that you've defined. The function allows for the 3rd argument to have more dimensions, for instance scattered locations all measured at a set of times, but it is fine for the first 3 arguments to all be 1-D.
You're treating longitude only as a mask, that is keeping longitudes in the range between 35 and 120 degrees east, correct? So the result will not depend on longitude.
Since z is a depth, you should define your axis zax with /depth, that is positive values of depth increasing downwards. You don't say what error you're running into. What plot commands are you trying and what happens?
Ansley
On 5/4/2018 8:03 AM, Pearse Buchanan wrote:
Hello all,
I’m trying to interpolate observations of NO3 using the function “scat2gridgauss_yz”.
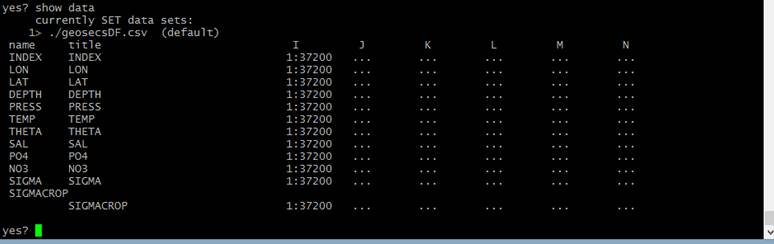
I successfully read in my data, which is structured as 1D column data, which looks like so:
As you can see, I have 37,200 observations.
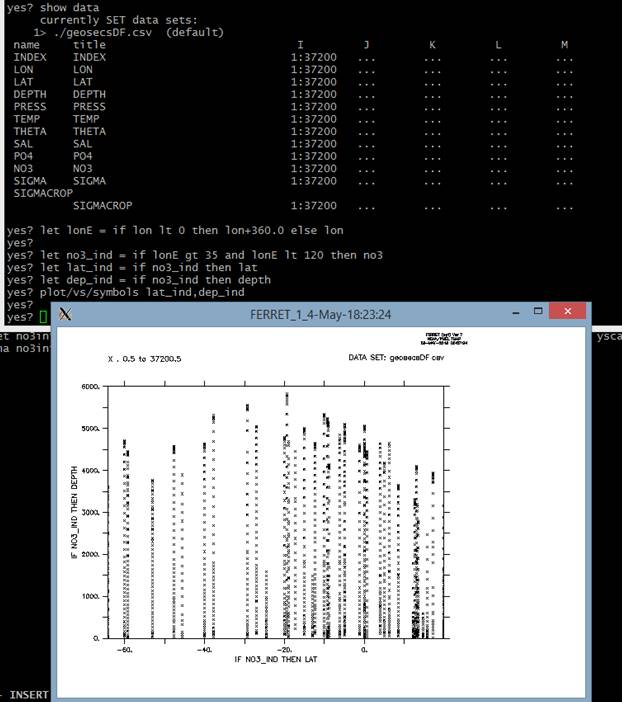
I now want to select my data from the Indian Ocean, where I eventually would like to do my interpolation using scat2gridgauss_yz.
Plotting the locations it seems like this has worked nicely.
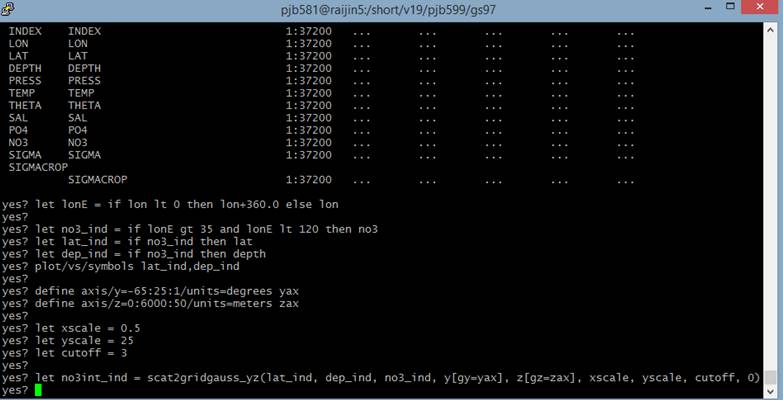
However, the problem then arises when I want to use “scat2gridgauss_yz” to interpolate this data. My command is:
And when trying to plot the data, I have an error. This error is related (I think!) to the fact that my NO3 data is in a 1D form. That is, it doesn’t have y,z coordinates. Scat2gridgauss requires that the variable in the 3rd argument be of 2 dimensions. Specifically, it must relate to the 2 dimensions of the 1st and 2nd arguments, in this case Latitude and Depth.
My question, therefore, is how to place my 1D NO3 data onto 2 dimensions of the lat,depth coordinates.
Thanks all,
Pearse
Pearse J. Buchanan
Institute for Marine and Antarctic Studies (IMAS), University of Tasmania
PhD Candidate / CSIRO-UTAS Quantitative Marine Science
ARC Centre of Excellence in Climate System Science – UTAS student representative
University of Tasmania Electronic Communications Policy (December, 2014).
This email is confidential, and is for the intended recipient only. Access, disclosure, copying, distribution, or reliance on any of it by anyone outside the intended recipient organisation is prohibited and may be a criminal offence. Please delete if obtained in error and email confirmation to the sender. The views expressed in this email are not necessarily the views of the University of Tasmania, unless clearly intended otherwise.